Thursday, September 6, 2012
Study Notes3 Predictive Modeling with Logistic Regression: group redundant variables
Sunday, September 2, 2012
Study Notes2 Predictive Modeling with Logistic Regression: collapse levels of contingency table
/* In the contingency table if there are many levels for a variable, then at some levels */
/* of this variable there may be no events(or all events) happening. This is called quasi */
/* completion problem(all separation). If this happens, the MLE will not exist since the */
/* regression coefficient will be infinit if want to maximize the likelihood */
/* A good solution is to collapse(cluster) some levels for the given categorical variable */
/*Variable Cluster_code has many levels which cause 0 events at some levels of cluster_code*/
/*This is called Quasi-completion problem*/
proc means data=pva1 nway missing;
class cluster_code;
var target_b;
output out=level mean=prop;
run;
proc print data=level width=min;
run;
ods output clusterhistory=cluster;
/* method=ward will use Greenacre's method to collapse levels of contingency tables */
/* the levels are clusterd based on the redunction in the chisq test of association */
/* levels with similar marginal response rate will be merged */
/* levels with least chisq value decreased will be merged, this is from the */
/* Semipartial R-Square value of the proc cluster output */
proc cluster data=level method=ward outtree=fortree plots=(dendrogram(vertical height=rsq));
freq _freq_;
var prop;
id cluster_code;
run;
proc freq data=pva1;
tables cluster_code*target_b / chisq;
output out=chi(keep=_pchi_) chisq;
run;
/* _pchi_ = 112.009 */
/* # of levels final is the one that giving least log(p_value) */
data cutoff;
if _n_ = 1 then set chi;
set cluster;
chisquare=_pchi_*rsquared;
degfree=numberofclusters-1;
logpvalue=logsdf('CHISQ',chisquare,degfree);
run;
proc sgplot data=cutoff;
scatter y=logpvalue x=numberofclusters / markerattrs=(color=blue symbol=circlefilled);
xaxis label="Number of Clusters";
yaxis label="Log of p-value";
title "Plot of the Log of p-value by Number of Clusters";
run;
/* number of clusters = 5 having the minimum logpvalue */
proc sql;
select numberofclusters into :ncl
from cutoff
having logpvalue=min(logpvalue);
quit;
ods html close;
proc tree data=fortree nclusters=&ncl out=clus;
id cluster_code;
run;
ods html;
proc sort data=clus;
by clusname;
run;
data clus_fmt;
retain fmtname "clus_fmt";
set clus;
start=cluster_code;
end=cluster_code;
label=clusname;
run;
/* at last use proc format to format the variable levels into clusters */
/* The formated fmt_cluster_code can be used as a categorical variable */
proc format cntlin=clus_fmt;
select clus_fmt;
run;
data pva1;
set pva1;
fmt_cluster_code=put(cluster_code,clus_fmt.);
run;
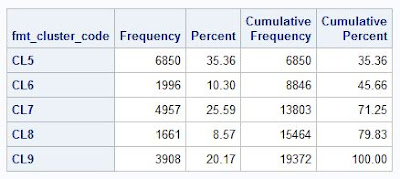
proc freq data=pva1;
table fmt_cluster_code / missing list;
run;

Saturday, September 1, 2012
Study Notes1 Predictive Modeling with Logistic Regression: impute missing data with proc stdize for continuous variable
/*This is the study notes from SAS online training. It includes why use Logistic Regression, how to clean the data */
/*(impute missing value, cluster rare events levels, variable clustering, variable screening), how to build */
/*logistic regression and how to measure the performance of the model */
libname mydata "D:\SkyDrive\sas_temp";
proc datasets lib=mydata;
contents data=_all_;
run;
*** How to impute the missing data by proc stdize ***;
/*data preparation*/
data pva(drop=control_number);
set mydata.pva_raw_data;
run;
/*use proc means with nmiss to check how many obs are missing for each variable*/
proc means data=pva nmiss min max median;
var donor_age income_group wealth_rating;
run;
/*use array to set indicator for missing obs, if missing then indicatd as 1*/
data pva;
set pva;
array a_mi{*} mi_donor_age mi_income_group mi_wealth_rating;
array a_var{*} donor_age income_group wealth_rating;
do i=1 to dim(a_mi);
a_mi(i)=(a_var(i)=.);
end;
run;
/*group data into 3 groups by recent_response_prop from low to high: first 1/3 grp_resp is 0, next 1/3 of data is 1 */
/*the last 1/3 is 2. In the same way for grp_amt. So totally there are 9 groups of data considering grp_resp and grp_ant */
/*obs numbers in grp_resp level or grp_amt level are similar, but at grp_resp*grp_amt level is different */
proc rank data=pva out=pva groups=3;
var recent_response_prop recent_avg_gift_amt;
ranks grp_resp grp_amt;
run;
proc freq data=pva;
table grp_resp grp_amt / missing list;
table grp_resp*grp_amt / missing list;
run;
/*sort the data by grp_resp and grp_amt*/
proc sort data=pva;
by grp_resp grp_amt;
run;
/*impute the missing data in each group formed by grp_resp*grp_amt, by the median value of non-missing data in that group*/
/*after the imputation, the data will have imputed value as well as the missing value indicator */
proc stdize data=pva method=median reponly out=pva1;
by grp_resp grp_amt;
var donor_age income_group wealth_rating;
run;
/*check the imputed value in each group*/
proc means data=pva median;
class grp_resp grp_amt;
var donor_age income_group wealth_rating;
run;

/*there are some other ways to impute, like cluster imputation using proc fastclus */
/*or using EM, MCMC, Regression to impute in proc mi(see UCLA ATS) */
/*proc mi can also use logistic regression to impute the categorical variables */
/* http://www.ats.ucla.edu/stat/sas/seminars/missing_data/part1.htm */
/* http://www.ats.ucla.edu/stat/sas/seminars/missing_data/part2.htm */