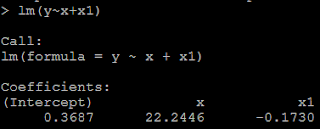Suppose U1~uniform(0,1), U2~uniform(0,1), and U1 is independent of U2, then what is the distribution of U1-U2?
The solution is in the picture attached. Then based on this, what is the distribution of U1+U2? The density should be of the same shape while it moves 1 unit to the right. The reason is: if U2 is uniform, then U3=1-U2 is uniform. so U1+U2 is the same as U1+U2=U1-U3+1. So it's density is the same as U1-U2 with i unit right transfer.
The answer can be verified in R. Density plot in R:
u1=runif(1000000,0,1) u2=runif(1000000,0,1) # Z is the difference of the two uniform distributed variable z=u1-u2 plot(density(z), main="Density Plot of the Difference of Two Uniform Variable", col=3) # X is the summation of the two uniform distributed variable x=u1+u2 plot(density(x), main="Density Plot of the Summation of Two Uniform Variable", col=3)